Researcher Profiles

Asiri Ediriwickrema, M.D., M.H.S
2023 Funding recipient
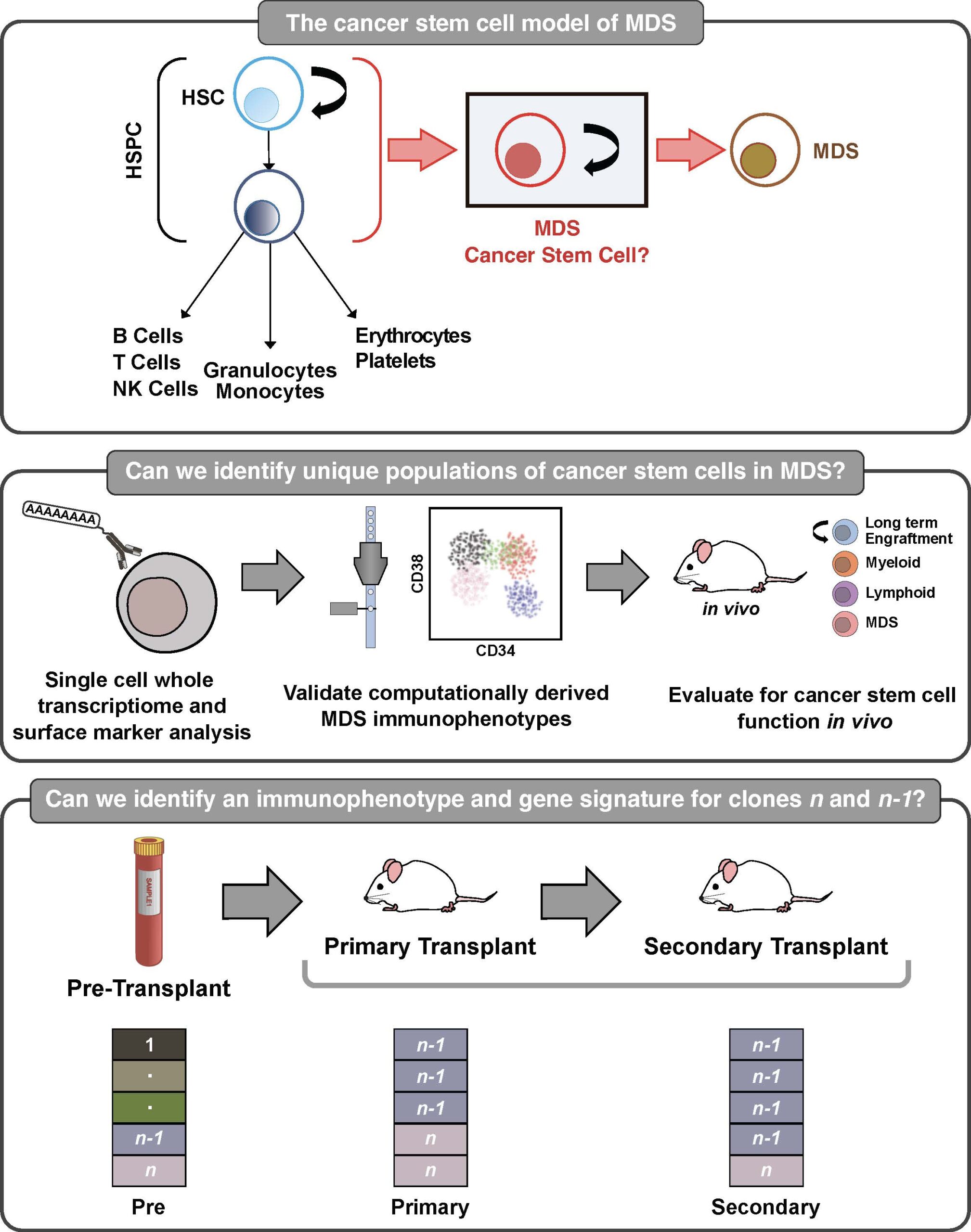
Characterizing high risk cancer stem cells in myelodysplastic neoplasms
EvansMDS Young Investigator Award 2023
PROJECT SUMMARY
Myelodysplastic neoplasms (MDS) are aggressive and often fatal cancers arising from the bone marrow. Despite aggressive chemotherapy and bone marrow transplant, patients with high-risk disease often die from treatment resistance, which is reflected by a poor average survival rate of 9 months. Several studies suggest that MDS is organized in a cellular hierarchy with a rare, self-renewing, cancer stem cell (CSC) giving rise to the remaining cancer progeny. The MDS-CSC is thought to initiate and maintain MDS, and therefore needs to be eliminated in order to cure patients. Unfortunately, CSCs are poorly defined, which has limited the ability to directly connect CSCs with worse clinical outcomes in MDS. It is not known if CSCs are prevalent in the majority of these patients, and if they express specific surface markers that can be used for disease monitoring and/or as therapeutic targets. Moreover, the biological mechanisms for CSC pathogenesis are unknown which further limits our understanding of their clinical relevance. There is a critical need to first improve our methods for isolating and studying these clinically relevant cells in MDS. My long-term goal is to improve MDS outcomes by developing cell-specific monitoring strategies and patient-specific anti-MDS therapies. To address this goal, the next step and overall objective of this application is to dissect the cellular heterogeneity of MDS through multiomic single cell analysis and rigorous functional studies. The central hypothesis is that MDS cells that co-express high-risk genes are not only distinct CSCs that can be purified and studied using specific surface makers, but are also the primary cells within this heterogenous subpopulation that contributes to poor outcomes. The rationale for the proposed research is that eradicating MDS-CSCs is necessary for curing patients. Guided by strong preliminary analysis, the central hypothesis will be tested by pursuing two specific aims that rigorously dissect the cellular heterogeneity of MDS using multiomic single cell analyses in order to design a surface marker panel to isolate and study MDS-CSCs. The overall contribution is expected to be the development of an actionable CSC immunophenotyping panel, and discovery of a newly defined, clinically relevant CSC-enriched sub-population(s) in human MDS. This contribution will be significant because it will allow us to not only improve patient outcomes by optimizing disease monitoring strategies but it will also facilitate the design of curative, patient-specific therapies in MDS.